First lines of code:
class TROPOMI(object):
# Installs geemap package
fme.exe python -m pip install geemap --target [x]
import fme
import fmeobjects
import ee
import geemap
import os
First lines of code:
class TROPOMI(object):
# Installs geemap package
fme.exe python -m pip install geemap --target [x]
import fme
import fmeobjects
import ee
import geemap
import os
First of all, line 5 isn't valid Python. You cannot execute fme.exe like that.
Secondly, when using fme.exe to install Python packages, make sure that you install for the same version of the Python interpreter that you'll be using in your workspace. You can force a specific Python version by appending either FME_PYTHON_VERSION 27 or FME_PYTHON_VERSION 37 at the end of the command, e.g.
fme.exe python -m pip install geemap --target [x] FME_PYTHON_VERSION 37
First of all, line 5 isn't valid Python. You cannot execute fme.exe like that.
Secondly, when using fme.exe to install Python packages, make sure that you install for the same version of the Python interpreter that you'll be using in your workspace. You can force a specific Python version by appending either FME_PYTHON_VERSION 27 or FME_PYTHON_VERSION 37 at the end of the command, e.g.
fme.exe python -m pip install geemap --target [x] FME_PYTHON_VERSION 37
Ah, I mistook command shell code as Python code. I now ran this:
fme.exe python -m pip install geemap --target [x] FME_PYTHON_VERSION 38in cmd and it seems to have installed the necessary dependencies.
How can I refer to [x] when trying to run this Python Script in FME?
import ee
import geemap
import os
ee.Initialize()
countries = ee.FeatureCollection("FAO/GAUL/2015/level0")
NL = countries.filterMetadata('ADM0_NAME', 'equals', 'Netherlands')
image = ee.ImageCollection('COPERNICUS/S5P/OFFL/L3_NO2') \
.select('tropospheric_NO2_column_number_density') \
.filterDate('2019-06-02', '2019-06-03') \
.mean()
roi = NL.geometry()
out_dir = os.path.join(os.path.expanduser('~'), 'Downloads')
filename = os.path.join(out_dir, 'NO2.tif')
image = image.clip(roi).unmask()
geemap.ee_export_image(image, filename=filename, scale=2000, region=roi, file_per_band=False)
Ah, I mistook command shell code as Python code. I now ran this:
fme.exe python -m pip install geemap --target [x] FME_PYTHON_VERSION 38in cmd and it seems to have installed the necessary dependencies.
How can I refer to [x] when trying to run this Python Script in FME?
import ee
import geemap
import os
ee.Initialize()
countries = ee.FeatureCollection("FAO/GAUL/2015/level0")
NL = countries.filterMetadata('ADM0_NAME', 'equals', 'Netherlands')
image = ee.ImageCollection('COPERNICUS/S5P/OFFL/L3_NO2') \
.select('tropospheric_NO2_column_number_density') \
.filterDate('2019-06-02', '2019-06-03') \
.mean()
roi = NL.geometry()
out_dir = os.path.join(os.path.expanduser('~'), 'Downloads')
filename = os.path.join(out_dir, 'NO2.tif')
image = image.clip(roi).unmask()
geemap.ee_export_image(image, filename=filename, scale=2000, region=roi, file_per_band=False)
If you defined [x] correctly when installing the module, FME should take care of the rest for you, and you should not need to refer to it anymore.
Ah, I mistook command shell code as Python code. I now ran this:
fme.exe python -m pip install geemap --target [x] FME_PYTHON_VERSION 38in cmd and it seems to have installed the necessary dependencies.
How can I refer to [x] when trying to run this Python Script in FME?
import ee
import geemap
import os
ee.Initialize()
countries = ee.FeatureCollection("FAO/GAUL/2015/level0")
NL = countries.filterMetadata('ADM0_NAME', 'equals', 'Netherlands')
image = ee.ImageCollection('COPERNICUS/S5P/OFFL/L3_NO2') \
.select('tropospheric_NO2_column_number_density') \
.filterDate('2019-06-02', '2019-06-03') \
.mean()
roi = NL.geometry()
out_dir = os.path.join(os.path.expanduser('~'), 'Downloads')
filename = os.path.join(out_dir, 'NO2.tif')
image = image.clip(roi).unmask()
geemap.ee_export_image(image, filename=filename, scale=2000, region=roi, file_per_band=False)
Thanks for your help so far David :)
Not sure how FME knows where [x] is, seeing as it is a custom folder I specified in the cmd install. I now attempt to install the package in FME like so:
try:
import geemap
except ImportError:
print("Installing geemap ...")
subprocess.check_call(["fme.exe", "python", "-m", "pip", "install", "geemap", "--target", "Q:\K\EIN\BU500\Algemeen\Afstudeer\tropomi\geemap"])When I run the script above, I get the following error...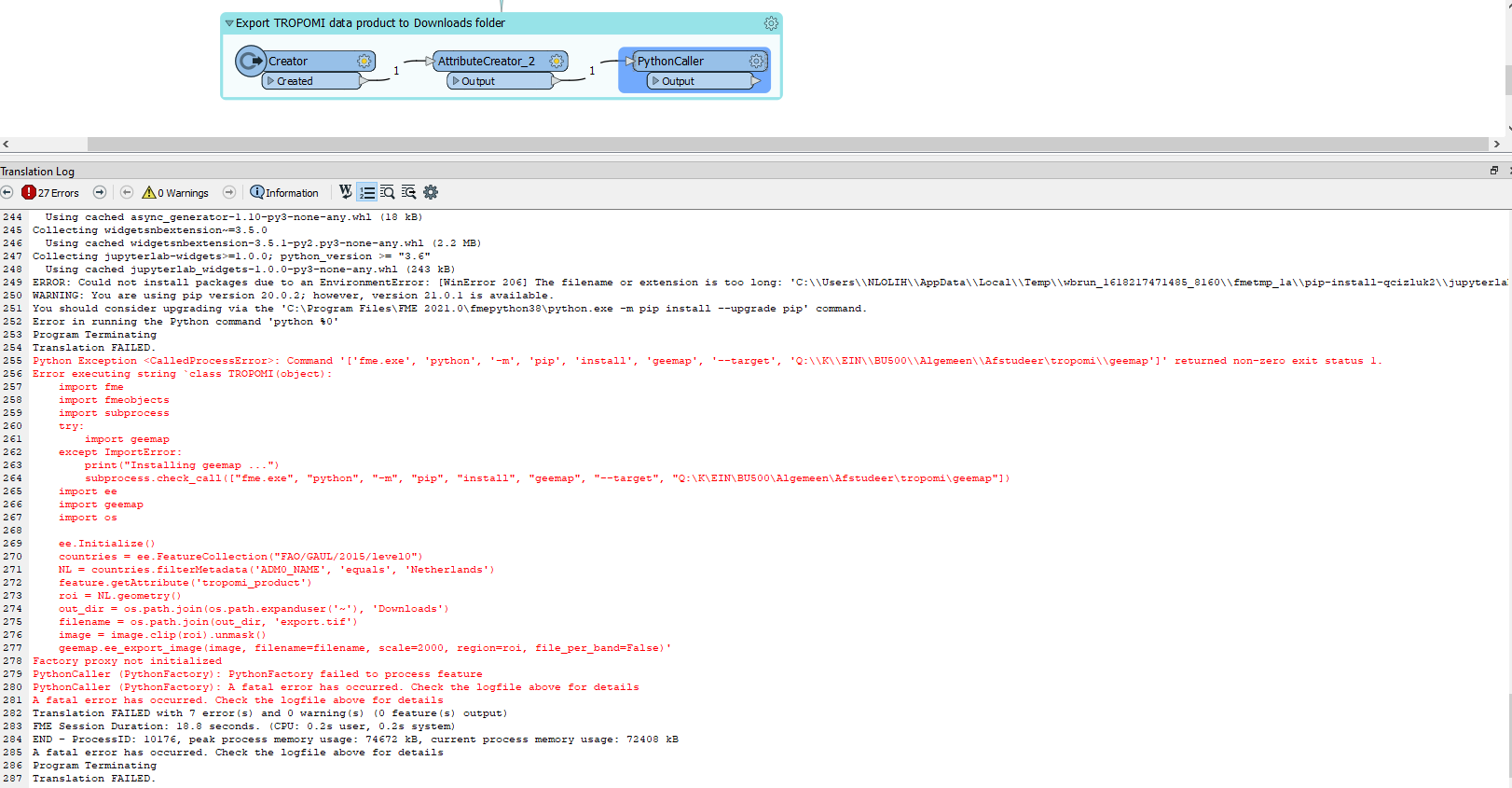
Ah, I mistook command shell code as Python code. I now ran this:
fme.exe python -m pip install geemap --target [x] FME_PYTHON_VERSION 38in cmd and it seems to have installed the necessary dependencies.
How can I refer to [x] when trying to run this Python Script in FME?
import ee
import geemap
import os
ee.Initialize()
countries = ee.FeatureCollection("FAO/GAUL/2015/level0")
NL = countries.filterMetadata('ADM0_NAME', 'equals', 'Netherlands')
image = ee.ImageCollection('COPERNICUS/S5P/OFFL/L3_NO2') \
.select('tropospheric_NO2_column_number_density') \
.filterDate('2019-06-02', '2019-06-03') \
.mean()
roi = NL.geometry()
out_dir = os.path.join(os.path.expanduser('~'), 'Downloads')
filename = os.path.join(out_dir, 'NO2.tif')
image = image.clip(roi).unmask()
geemap.ee_export_image(image, filename=filename, scale=2000, region=roi, file_per_band=False)
[x] should refer to a folder that is known to FME, e.g.
C:\Users\<username>\Documents\FME\Plugins\Python\python38 for Python 3.8 modules.
Ah, I mistook command shell code as Python code. I now ran this:
fme.exe python -m pip install geemap --target [x] FME_PYTHON_VERSION 38in cmd and it seems to have installed the necessary dependencies.
How can I refer to [x] when trying to run this Python Script in FME?
import ee
import geemap
import os
ee.Initialize()
countries = ee.FeatureCollection("FAO/GAUL/2015/level0")
NL = countries.filterMetadata('ADM0_NAME', 'equals', 'Netherlands')
image = ee.ImageCollection('COPERNICUS/S5P/OFFL/L3_NO2') \
.select('tropospheric_NO2_column_number_density') \
.filterDate('2019-06-02', '2019-06-03') \
.mean()
roi = NL.geometry()
out_dir = os.path.join(os.path.expanduser('~'), 'Downloads')
filename = os.path.join(out_dir, 'NO2.tif')
image = image.clip(roi).unmask()
geemap.ee_export_image(image, filename=filename, scale=2000, region=roi, file_per_band=False)
Yes, that link is where I got the code in my first post, but I am trying to install it to a folder accessible by my network. I need this workspace to run from any PC that has access to [x], is this possible? I am trying to avoid everyone having to install the package to their own PC.
Ah, I mistook command shell code as Python code. I now ran this:
fme.exe python -m pip install geemap --target [x] FME_PYTHON_VERSION 38in cmd and it seems to have installed the necessary dependencies.
How can I refer to [x] when trying to run this Python Script in FME?
import ee
import geemap
import os
ee.Initialize()
countries = ee.FeatureCollection("FAO/GAUL/2015/level0")
NL = countries.filterMetadata('ADM0_NAME', 'equals', 'Netherlands')
image = ee.ImageCollection('COPERNICUS/S5P/OFFL/L3_NO2') \
.select('tropospheric_NO2_column_number_density') \
.filterDate('2019-06-02', '2019-06-03') \
.mean()
roi = NL.geometry()
out_dir = os.path.join(os.path.expanduser('~'), 'Downloads')
filename = os.path.join(out_dir, 'NO2.tif')
image = image.clip(roi).unmask()
geemap.ee_export_image(image, filename=filename, scale=2000, region=roi, file_per_band=False)
It should be doable, if you add something like the following at the top of every Python code block that imports the "geemap" module, before the "import geemap" statement:
import sys
geemap_path = "[x]"
if not geemap_path in sys.path:
sys.path.append(geemap_path)Basically it adds [x] to the Python search path, if not already present.
Ah, I mistook command shell code as Python code. I now ran this:
fme.exe python -m pip install geemap --target [x] FME_PYTHON_VERSION 38in cmd and it seems to have installed the necessary dependencies.
How can I refer to [x] when trying to run this Python Script in FME?
import ee
import geemap
import os
ee.Initialize()
countries = ee.FeatureCollection("FAO/GAUL/2015/level0")
NL = countries.filterMetadata('ADM0_NAME', 'equals', 'Netherlands')
image = ee.ImageCollection('COPERNICUS/S5P/OFFL/L3_NO2') \
.select('tropospheric_NO2_column_number_density') \
.filterDate('2019-06-02', '2019-06-03') \
.mean()
roi = NL.geometry()
out_dir = os.path.join(os.path.expanduser('~'), 'Downloads')
filename = os.path.join(out_dir, 'NO2.tif')
image = image.clip(roi).unmask()
geemap.ee_export_image(image, filename=filename, scale=2000, region=roi, file_per_band=False)
Hmm, I tried this but got this error. Which hints at an incomplete geemap install (?)
Python Exception <ModuleNotFoundError>: No module named 'ee'How can I best install the geemap package to [x] within the script in FME? Is there anyway to do this?
Ah, I mistook command shell code as Python code. I now ran this:
fme.exe python -m pip install geemap --target [x] FME_PYTHON_VERSION 38in cmd and it seems to have installed the necessary dependencies.
How can I refer to [x] when trying to run this Python Script in FME?
import ee
import geemap
import os
ee.Initialize()
countries = ee.FeatureCollection("FAO/GAUL/2015/level0")
NL = countries.filterMetadata('ADM0_NAME', 'equals', 'Netherlands')
image = ee.ImageCollection('COPERNICUS/S5P/OFFL/L3_NO2') \
.select('tropospheric_NO2_column_number_density') \
.filterDate('2019-06-02', '2019-06-03') \
.mean()
roi = NL.geometry()
out_dir = os.path.join(os.path.expanduser('~'), 'Downloads')
filename = os.path.join(out_dir, 'NO2.tif')
image = image.clip(roi).unmask()
geemap.ee_export_image(image, filename=filename, scale=2000, region=roi, file_per_band=False)
Did you watch out for the \ character?
Either it needs to be doubled, e.g.
geemap_path = "c:\\temp\\modules\\geemap"Or you'll need to tell Python that it's a raw string by prefixing it by a lower case "r", e.g.
geemap_path = r"c:\temp\modules\geemap"
Ah, I mistook command shell code as Python code. I now ran this:
fme.exe python -m pip install geemap --target [x] FME_PYTHON_VERSION 38in cmd and it seems to have installed the necessary dependencies.
How can I refer to [x] when trying to run this Python Script in FME?
import ee
import geemap
import os
ee.Initialize()
countries = ee.FeatureCollection("FAO/GAUL/2015/level0")
NL = countries.filterMetadata('ADM0_NAME', 'equals', 'Netherlands')
image = ee.ImageCollection('COPERNICUS/S5P/OFFL/L3_NO2') \
.select('tropospheric_NO2_column_number_density') \
.filterDate('2019-06-02', '2019-06-03') \
.mean()
roi = NL.geometry()
out_dir = os.path.join(os.path.expanduser('~'), 'Downloads')
filename = os.path.join(out_dir, 'NO2.tif')
image = image.clip(roi).unmask()
geemap.ee_export_image(image, filename=filename, scale=2000, region=roi, file_per_band=False)
Good point. I have added the double backslash.
If I run the code below, I get the following error:
Python Exception <ImportError>: cannot import name 'Box' from 'box' (C:\Program Files\FME\python\python38\box\__init__.pyc)Have you seen this error before?
The code I am now running:
class TROPOMI(object):
import sys
geemap_path = "C:\\Users\\ollih\\Desktop\\geemap"
if not geemap_path in sys.path:
sys.path.append(geemap_path)
import fme
import fmeobjects
import ee
import geemap
import os
ee.Initialize()
countries = ee.FeatureCollection("FAO/GAUL/2015/level0")
NL = countries.filterMetadata('ADM0_NAME', 'equals', 'Netherlands')
feature.getAttribute('tropomi_product')
roi = NL.geometry()
out_dir = os.path.join(os.path.expanduser('~'), 'Downloads')
filename = os.path.join(out_dir, 'export.tif')
image = image.clip(roi).unmask()
geemap.ee_export_image(image, filename=filename, scale=2000, region=roi, file_per_band=False)
Ah, I mistook command shell code as Python code. I now ran this:
fme.exe python -m pip install geemap --target [x] FME_PYTHON_VERSION 38in cmd and it seems to have installed the necessary dependencies.
How can I refer to [x] when trying to run this Python Script in FME?
import ee
import geemap
import os
ee.Initialize()
countries = ee.FeatureCollection("FAO/GAUL/2015/level0")
NL = countries.filterMetadata('ADM0_NAME', 'equals', 'Netherlands')
image = ee.ImageCollection('COPERNICUS/S5P/OFFL/L3_NO2') \
.select('tropospheric_NO2_column_number_density') \
.filterDate('2019-06-02', '2019-06-03') \
.mean()
roi = NL.geometry()
out_dir = os.path.join(os.path.expanduser('~'), 'Downloads')
filename = os.path.join(out_dir, 'NO2.tif')
image = image.clip(roi).unmask()
geemap.ee_export_image(image, filename=filename, scale=2000, region=roi, file_per_band=False)
My first impression is a missing requirement or a dependency, or perhaps even some namespace problem. I can't really tell without further knowledge of the other modules.
Are you using a BoxConnector transformer somewhere in your workspace? The problem seems to be related to that.
Enter your username or e-mail address. We'll send you an e-mail with instructions to reset your password.